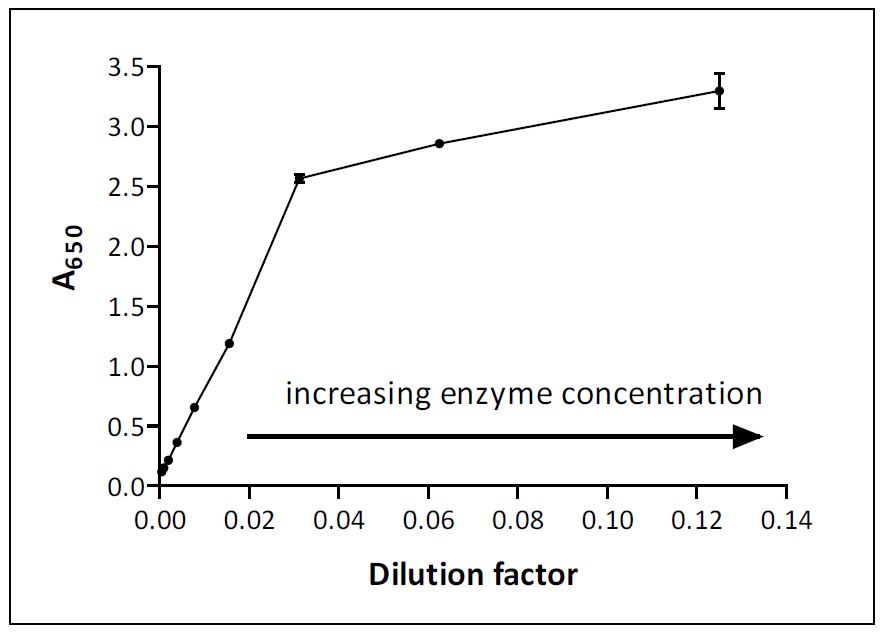
Guide to Enzyme Unit Definitions and Assay Design | Biomol Blog | Resources | Biomol GmbH - Life Science Shop

Automated relative binding free energy calculations: from SMILES to ΔΔG – BioExcel – Centre of Excellence for Computation Biomolecular Research
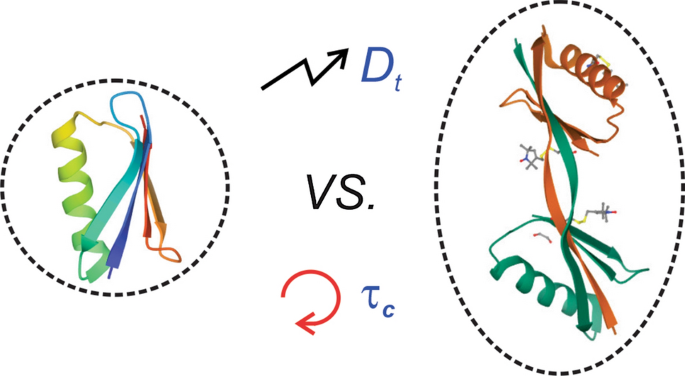
NMR measurement of biomolecular translational and rotational motion for evaluating changes of protein oligomeric state in solution | SpringerLink

L7-1 Slides courtesy of Prof M L Kraft, Chemical & Biomolecular Engr Dept, University of Illinois at Urbana-Champaign. Review: Liquid Phase Reaction in. - ppt download

Systematic discovery of biomolecular condensate-specific protein phosphorylation | Nature Chemical Biology

Optimized Magnesium Force Field Parameters for Biomolecular Simulations with Accurate Solvation, Ion-Binding, and Water-Exchange Properties in SPC/E, TIP3P-fb, TIP4P/2005, TIP4P-Ew, and TIP4P-D | Journal of Chemical Theory and Computation
CHARMM-GUI Free Energy Calculator for Absolute and Relative Ligand Solvation and Binding Free Energy Simulations
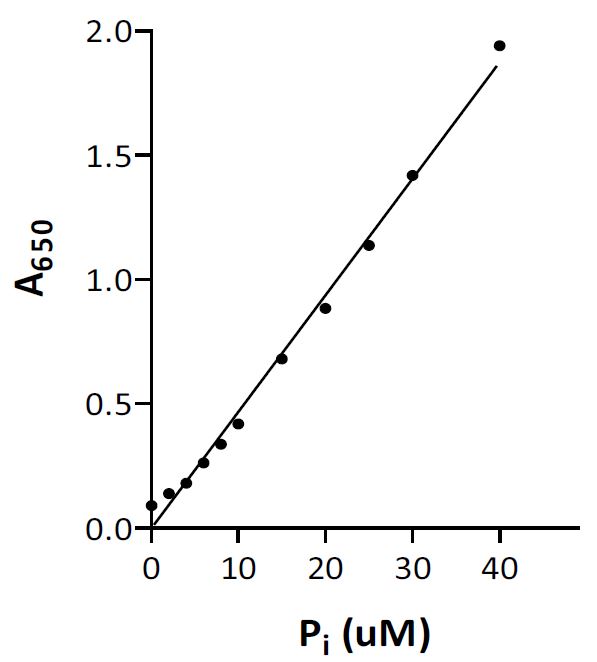
Guide to Enzyme Unit Definitions and Assay Design | Biomol Blog | Resources | Biomol GmbH - Life Science Shop

Artificial Biomolecular Channels: Enantioselective Transmembrane Transport of Amino Acids Mediated by Homochiral Zirconium Metal–Organic Cages | Journal of the American Chemical Society
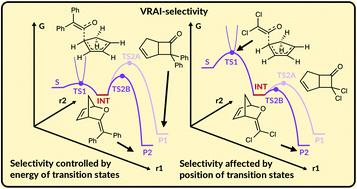
VRAI-selectivity: calculation of selectivity beyond transition state theory - Organic & Biomolecular Chemistry (RSC Publishing)
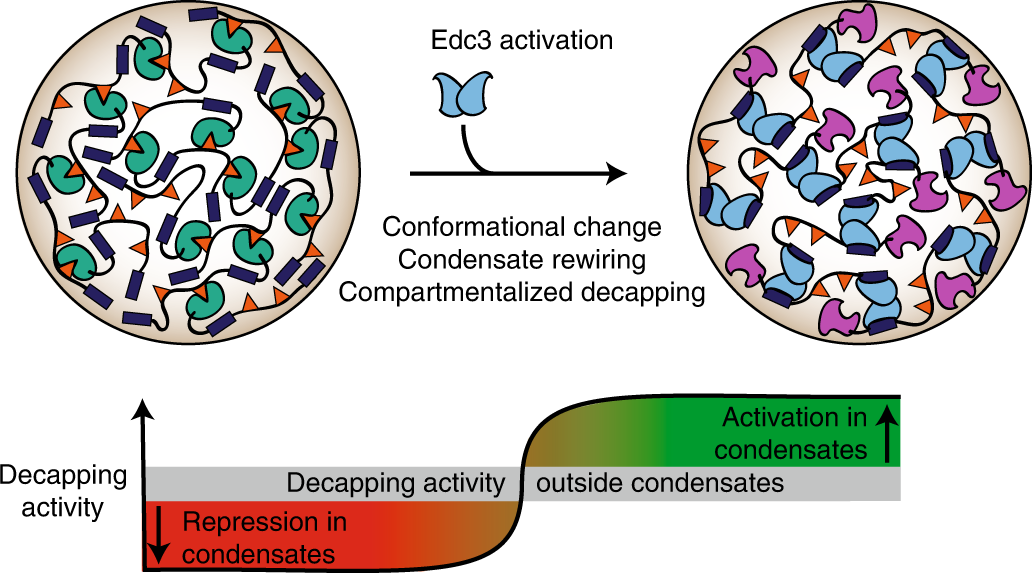
Biomolecular condensates amplify mRNA decapping by biasing enzyme conformation | Nature Chemical Biology

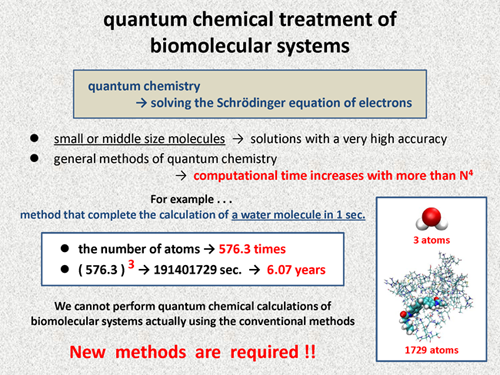
A Self-Improving Photosensitizer Discovery System via Bayesian Optimization and Quantum Chemical Calculation | Materials Chemistry | ChemRxiv | Cambridge Open Engage
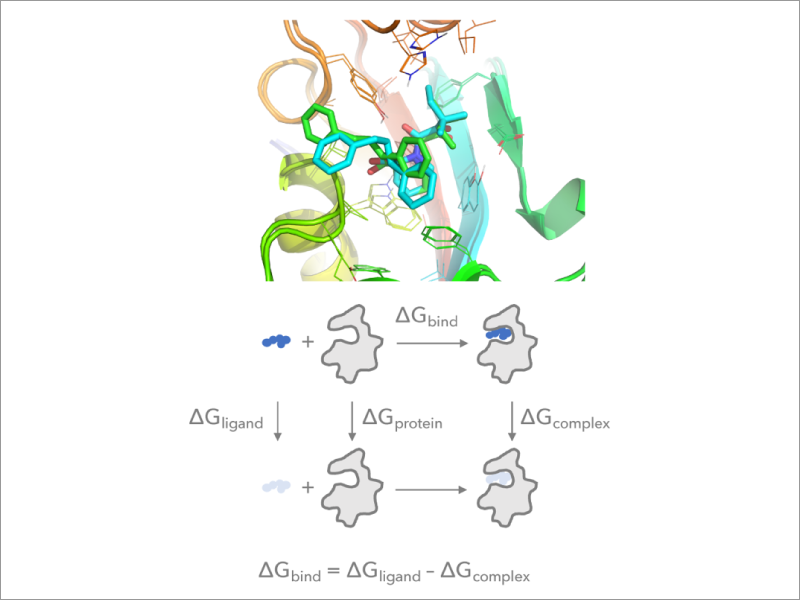
Development of free energy calculation methods for biomolecular associations – Laboratory for Biomolecular Function Simulation




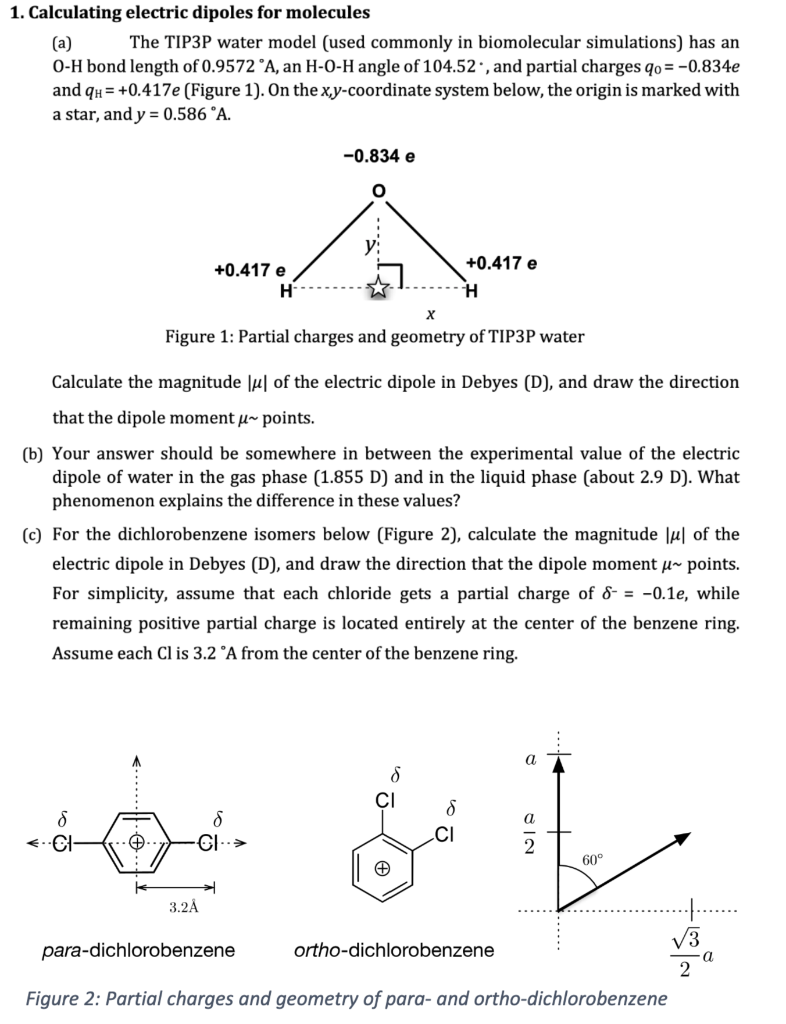



![Solved り(pM/min) 130 116 87 63 30 10 [S] (mole/liter) 65 x | Chegg.com Solved り(pM/min) 130 116 87 63 30 10 [S] (mole/liter) 65 x | Chegg.com](https://media.cheggcdn.com/media%2F451%2F451ce7ba-1a04-449b-9f54-e04ea89ffbe5%2Fimage)